The Challenge
Molecular phenotypes - transcripts, metabolites, accessible chromatin - shift in response to environment, development, and circadian rhythms. To detect meaningful associations in genetically diverse populations, tissue must be collected within a one-to-two hour window. The tighter the window, the stronger the signal.
Our Solution
Our lab first confronted this problem in 2020 as part of an ARPA-E sponsored research project. Jonathan Turkus was tasked with designing a system capable of sampling 750 corn plots in under two hours, with immediate flash-freezing in liquid nitrogen to prevent degradation of labile molecular phenotypes. The system he developed met that target, and years of iterative refinement have dramatically improved throughput. Our personal best, set in August 2021 during a sorghum diversity study, was 2,898 tubes collected from 966 plots in 92 minutes with a team of eleven samplers plus several coordinators managing hot-swapping of tube racks. That’s a sustained rate of one plot per minute (one tube every 20 seconds) per person for more than 1.5 hours.
Video
Full demonstration of the rapid field tissue collection system during a sampling campaign. Open on YouTube.
How It Works
At its core, the system combines pre-labeled tubes and leaf punches for sample collection with a carrier system that allows collectors to safely transport liquid nitrogen or dry ice through the field.
Trap-Door Cryogenic Carriers
Samples are dropped through a trap-door mechanism into liquid nitrogen or dry ice carried through the field. The design ensures that neither samples nor cryogen will leak out, even if the kit is dropped or a person trips, keeping both our people and our samples safe.
Spill-Proof Pre-Ordered Tube Trays
All tubes are pre-labeled and arranged in exact collection order before going to the field. Custom trays hold every tube securely and can be flipped 180 degrees without a single tube falling out - critical when one stumble could scatter hundreds of tubes and derail an entire experiment. Trays are hot-swappable: full trays are pre-positioned throughout the field, so samplers simply replace an empty tray with the next one and keep moving.

Photo
Pre-labeled tubes arranged in collection order and loaded into spill-proof trays.

Video
Spill-proof tray demonstration showing the rack fully inverted without tube loss.
Triplicate Parallel Sampling
Three biological samples are collected simultaneously from the same leaf or other tissue. The primary sample is used for analysis; if something goes wrong during extraction, QC, or data generation, backup samples collected from the same plant at the same moment are available. Because all samples from a population must come from the same time window on the same day, there is no opportunity to go back and re-collect a lost sample - backups are essential.
Alternatively, replicates can support entirely different assays - gene expression, metabolomics, and chromatin accessibility - from the same population, in the same field, at the same time.

Video
Demonstration of triplicate sample collection - three parallel biological samples taken simultaneously from the same tissue. View on Flickr.
Adoption
The system has been used by research groups across ten states and by multiple private companies for high-throughput molecular phenotyping under field conditions. We use it primarily for corn and sorghum, but other groups have successfully tested it with other crops.
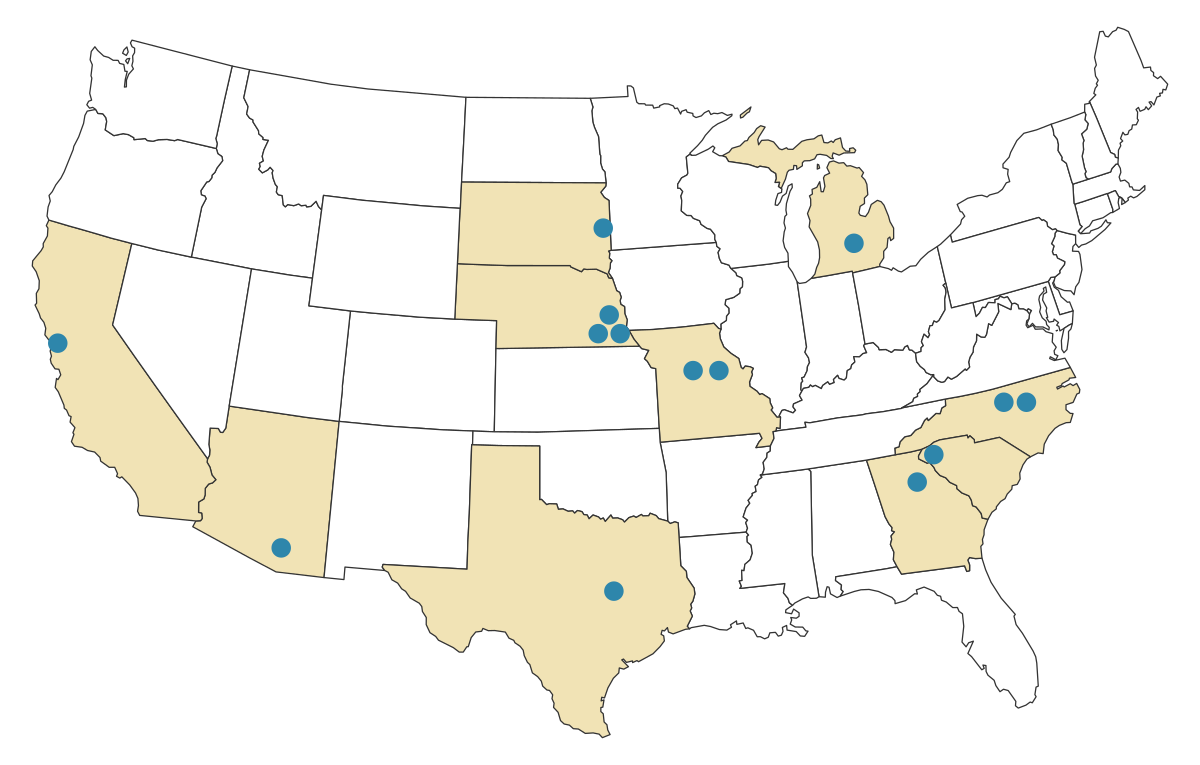
Research groups and companies using the rapid field tissue collection system.
Publications Using This System
- Grzybowski M, Schnable JC (2026). Haplotype-rich cis-regulation underlies transcriptomic diversity across the breeding history of maize (Zea mays). bioRxiv. doi: 10.64898/2026.02.19.706772. Preprint.
- Creach M, Webster B, Newton L, Turkus J, Schnable JC, Thompson AM, VanBuren R (2025). Predicting complex phenotypes using multi-omics data in maize. bioRxiv. doi: 10.1101/2025.09.30.679283. Preprint.
- Mathivanan RK, Pedersen C, Turkus J, Shrestha N, Ali W, Torres-Rodriguez JV, Mural RV, Obata T, Schnable JC (2025). Transcripts and genomic intervals associated with variation in metabolite abundance in maize leaves under field conditions. BMC Genomics. doi: 10.1186/s12864-025-11580-3.
- Mangal H, Linders K, Turkus J, Shrestha N, Long B, Kuang X, Cebert E, Torres-Rodriguez JV, Schnable JC (2025). Genes and pathways determining flowering time variation in temperate adapted sorghum. The Plant Journal. doi: 10.1111/tpj.70250.
- Torres-Rodriguez JV, Li D, Schnable JC (2025). Evolving best practices for transcriptome-wide association studies accelerate discovery of gene-phenotype links. Current Opinion in Plant Biology. doi: 10.1016/j.pbi.2024.102670.
- Torres-Rodriguez JV, Li D, Turkus J, Newton L, Davis J, Lopez-Corona L, Ali W, Sun G, Mural RV, Grzybowski M, Zamft B, Thompson AM, Schnable JC (2024). Population level gene expression can repeatedly link genes to functions in maize. The Plant Journal. doi: 10.1111/tpj.16801.
Contact / Collaborate
Interested in learning more or collaborating? Reach out to Jonathan Turkus or James Schnable.

Photo
The rapid field tissue collection system in use during a sorghum diversity panel sampling campaign. View on Flickr.
Acknowledgements
This work was supported in part by the Department of Energy’s Advanced Research Projects Agency-Energy (ARPA-E) under award DE-AR0001367, the Foundation for Food and Agriculture Research under Award No. 602757, and by the Nebraska Corn Checkoff Presidential Chair Endowment.
